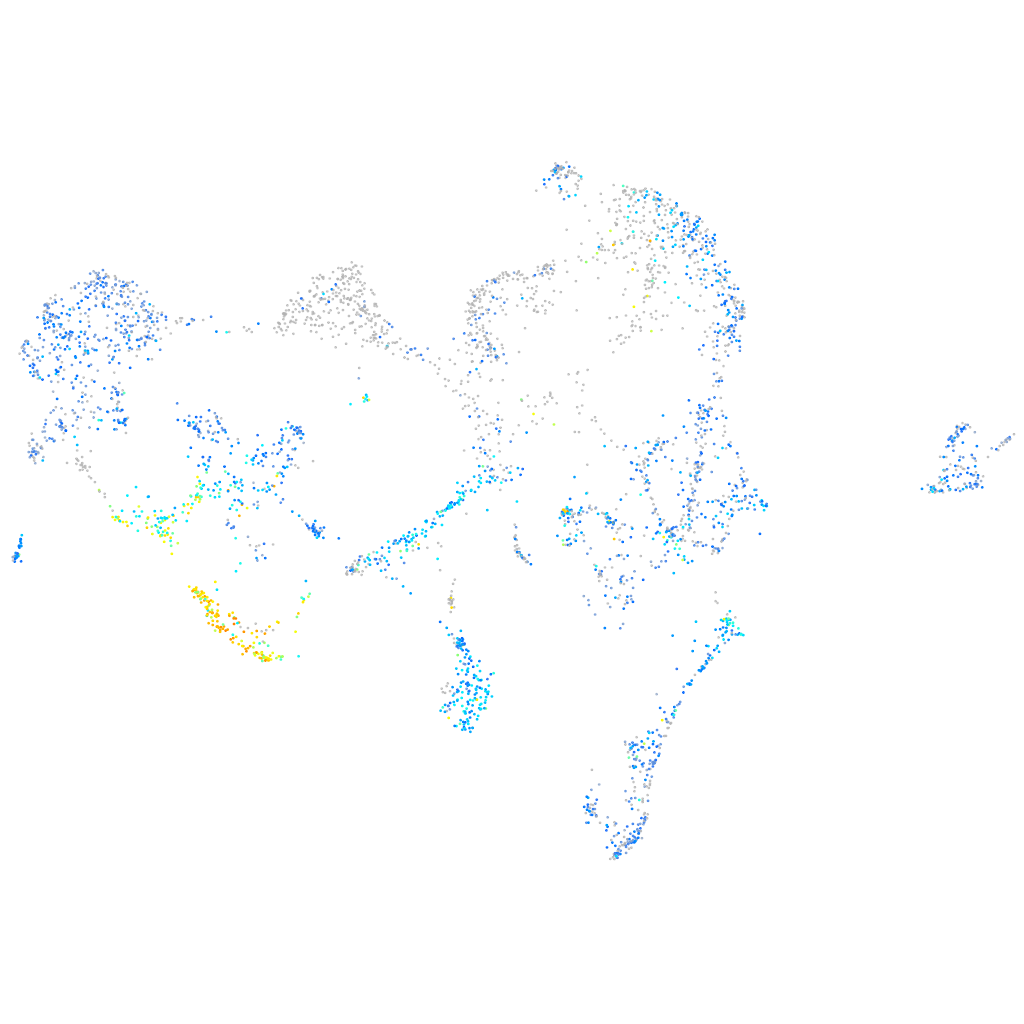
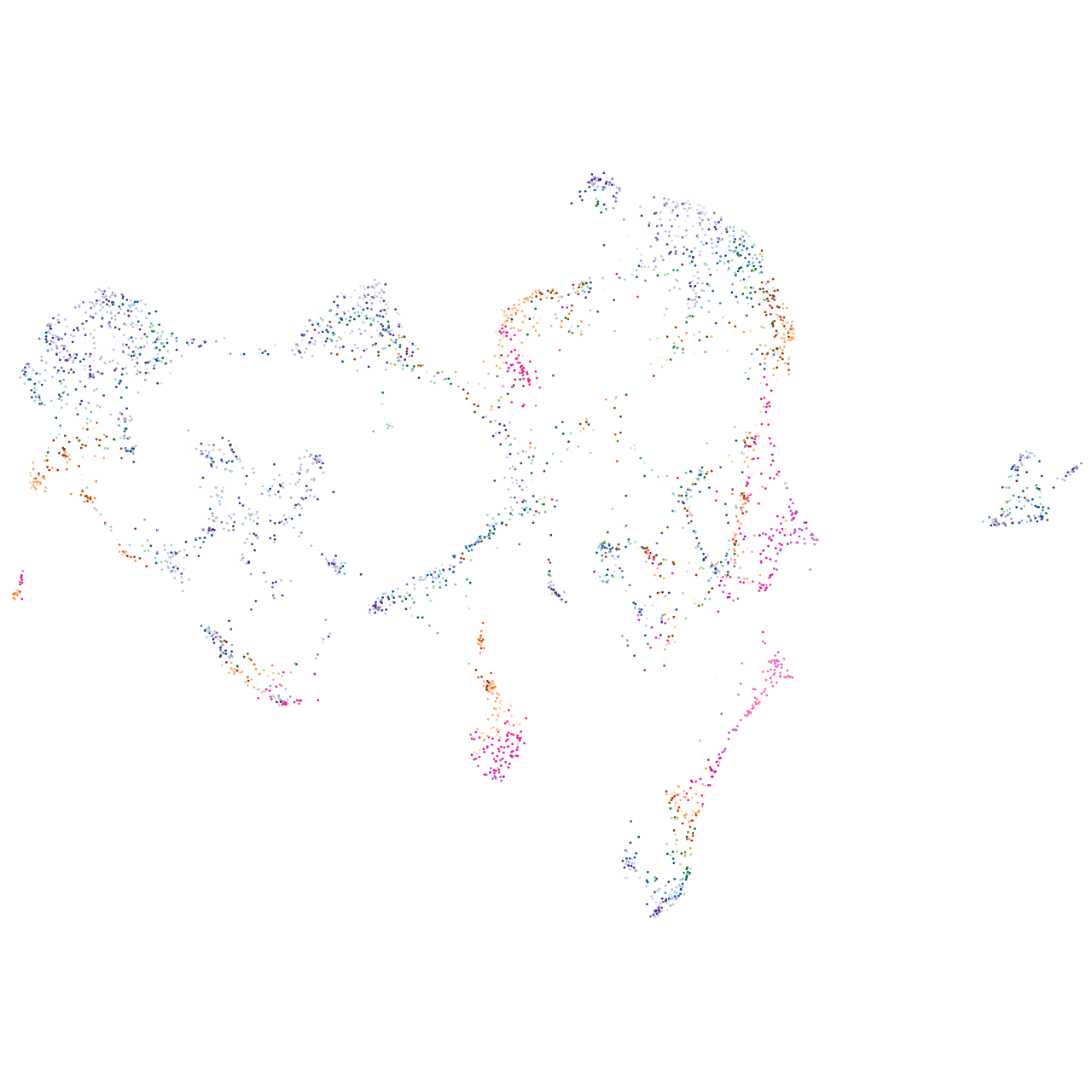
"ATPase Na+/K+ transporting subunit alpha 1a, tandem duplicate 1"
ZFIN 
Expression by stage/cluster


Correlated gene expression


| Positive correlation | Negative correlation | ||
| Gene | r | Gene | r |
| s100a11 | 0.665 | krt91 | -0.393 |
| trpv6 | 0.653 | icn | -0.314 |
| igfbp5a | 0.640 | si:dkey-193p11.2 | -0.308 |
| mafbb | 0.620 | atp1a1a.2 | -0.292 |
| FP016234.1 | 0.608 | malb | -0.283 |
| slc8a1b | 0.606 | LOC799574 | -0.247 |
| LOC101886621 | 0.602 | rgs5b | -0.239 |
| fthl29 | 0.590 | si:dkey-189h5.6 | -0.238 |
| fthl28 | 0.561 | si:ch73-359m17.9 | -0.231 |
| tpte | 0.549 | atp1a1a.3 | -0.227 |
| fthl30 | 0.543 | ndrg1a | -0.222 |
| fstb | 0.541 | ahnak | -0.216 |
| si:dkey-159a18.1 | 0.529 | slc12a10.2 | -0.215 |
| CABZ01102170.1 | 0.513 | prlra | -0.210 |
| fthl31 | 0.512 | sri | -0.207 |
| igf2b | 0.509 | lgals3b | -0.201 |
| pvalb8 | 0.500 | rnf183 | -0.200 |
| ankhb | 0.499 | clcn2c | -0.199 |
| mafba | 0.487 | nudt4a | -0.197 |
| usp2a | 0.475 | litaf | -0.196 |
| CU914776.1 | 0.474 | gsto1 | -0.196 |
| sgk2b | 0.469 | zgc:101785 | -0.196 |
| dscaml1 | 0.467 | agr2 | -0.196 |
| appb | 0.466 | nupr1b | -0.192 |
| vsnl1a | 0.451 | trim35-12 | -0.192 |
| camk2n1a | 0.451 | zgc:153284 | -0.190 |
| si:dkey-229e3.2 | 0.447 | zgc:163083 | -0.190 |
| ponzr5 | 0.423 | dap1b | -0.186 |
| dscama | 0.418 | nipal4 | -0.182 |
| gata3 | 0.412 | si:dkey-248g15.3 | -0.180 |
| pcdh17 | 0.411 | gcshb | -0.179 |
| gcm2 | 0.410 | s100z | -0.177 |
| gaa3-as | 0.407 | sdr16c5b | -0.176 |
| si:dkey-251i10.1 | 0.404 | pcdh11 | -0.174 |
| pappaa | 0.403 | tuft1a | -0.173 |